DNA Super Coiling:
Whether the DNA is single stranded or double stranded, inside the cell it is always subjected to bending, folding, over winding and under winding. Such changes make stable DNA into unstable or vice versa, because energy constraints. Thermodynamically complaint B-DNA contains 10.6 bp per turn and~35 A° length per turn and such E.coli DNA or a linear DNA without any constraints or stress or torsion, when placed on a flat surface it lays flat as a circular molecule or a linear molecule. Such DNA, which is free from stress, is called Relaxed DNA.
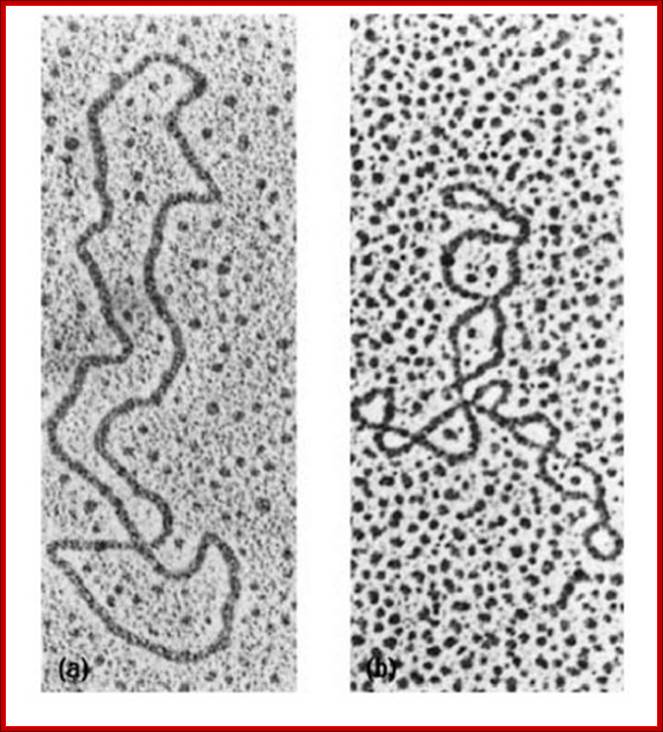
Relaxed circular and Supercoiled DNA; http://what-when-how.com/
This is an EM of one a linear DNA
Below-Circular and Supercoiled DNA
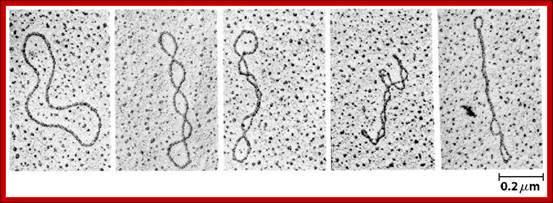
Electron micrographs of relaxed and supercoiled plasmid DNAs. The molecule at the left is relaxed, and the degree of supercoiling increases from left to right.
http://www.bioinfo.org.cn/
When a relaxed DNA is subjected to bends or openings of DNA, over winding or unwinding, its base pairs per turn changes, and the DNA is subjected to stress and strain. In order to overcome such distortion, which has rendered the DNA unstable, the DNA twist around itself , on its own axis, like a circular rubber band undergoing twisting; such a twists on its own thread is called Super Coiling; it is also referred to as tertiary structure, it can be positive or negative supercoiling.
![]()
This a linear relaxed DNA
This is relaxed circular DNA; http://en.wikibooks.org/
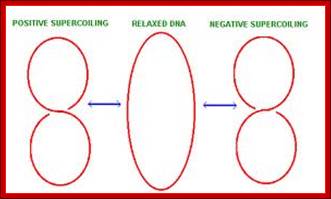
Positive super coiling:
To explain super coiling phenomenon, if a 5300 bp long circular DNA, which contains ~500 helical turns with 10.6bp per turn, if it is twisted 50 times by 360^° in right handed direction i.e. towards its own right handed coil, the DNA becomes over coiled or over winded and the helix becomes tight. It virtually means adding 50 more helical coils added to the already existing 500 helical turns, making the total number of turns to 550. So the 5300 bp long DNA has to accommodate 50 more turns, which is torsionally a stressed condition. This over winding is thermodynamically incompatible, so in order to accommodate the excess number of coils, the helical DNA twists on its own length in space in left handed mode. [ If a two jute fiber thread is over twisted, it coils on its own in opposite way]. If 5300 bp is divided by 550 turns, the number of base pairs per turn will be 9.6; it is a drastic deviation from the normal and thermodynamically feasible, relaxed and stable state of 10.6 per turn, like a rubber band twist, in left handed way, such a twist is called super coil; in this case the super coiling is positive. Such super coilings overcome stress energy and accommodate the extra 50 more helices or added coils to the existing 500 helical turns. It is because of super coiling; now, the number of base pairs per turn would be 10.6
Negative super coiling:
Similarly, if a 5300bp long circular right handed coiled DNA with 500° gets under wounded, and likely the DNA strands can separate. In this case the number of turns lost are 50 and to adjust the number of coils, the number of base pairs per turn will be11.6, which is again not thermodynamically feasible, hence the DNA takes a right handed twist on its own; the DNA helix is in the form of negatively super coiled DNA. This super coiling accommodates the stress energy and the DNA becomes stable only if the coil is restored to 10.6 bp per coil.
Generally the density of the super coils in a DNA is one super coil per 200bp or per ~20 turns of the helix. Negative super coils help in stabilizing certain DNA structures i.e. Z-DNA, cruciform DNA, and triplex DNA. It also helps in unwinding of the DNA during replication as replication bubble or during transcriptional initiation as transcriptional bubble.
Mechanism of super coiling:
One way in which DNA is regulated at a biophysical level is through supercoiling — positive or negative — meaning twisting up to a greater or lesser extent. The name “supercoiling” highlights the fact that this is “on top of” the molecule’s natural helical coil.
As with the stretching of other biomolecules, this action can expose binding sites (for transcription factors) and is crucial in regulating DNA replication, where negative supercoiling is required to allow processes such as replication, transcription and recombination to be carried out on the genome.
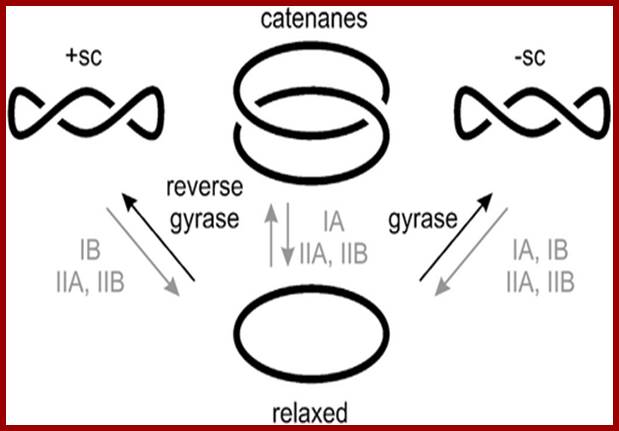
The catenane in the centre of the simple diagram here clearly requires a separation (by a topoisomerase) to get back to the circular, relaxed form. Likewise, the positive and negative values of supercoiling are determined by whether the gyrases apply tension with or counter to the turn of the α-helix. Negative supercoiling (-sc) favours strand separation, while +sc stabilises double-stranded DNA (dsDNA). http://biochemistri.es/
A cascade of DNA induced conformational changes prepares gyrase
for strand passage; http://biochemistri.es/
Coiling can only take place on a molecule that is either a closed loop (such as bacterial plasmids or DNA minicircles) or linear molecules that are fixed at certain points, such as by interacting proteins — making it for all intents and purposes the topological equivalent to a closed loop, and twist (or writhe) being converted to tension within the molecule rather than spinning motion.
The catenane in the center of the simple diagram here clearly requires a separation (by a topoisomerase) to get back to the circular, relaxed form. Likewise, the positive and negative values of supercoiling are determined by whether the gyrases apply tension with or counter to the turn of the α-helix. Negative supercoiling (-sc) favors strand separation, while +sc stabilizes double-stranded DNA (dsDNA).
Detection of super Coils:
Super coiled DNA can be detected by using gel electrophoresis. Super coiled DNA moves faster in an agarose gel than the relaxed DNA. If a purified plasmid DNA is run on a gel, and if one finds more than two bands, it means the one that moved faster is super coiled and the one that moves slower is nicked, so it moves slowly. By subjecting such super coiled DNA to Topoisomerases, an enzyme that nicks and relaxes the super coiled DNA, one cut per digestion and such digested DNA if it is run on an agarose gel, one can observe a number of bands from the bottom. The bottom most is the most super coiled and uncut. The band found next to the uncut shows that it is relaxed by one super coil. The uppermost band is the one, which is totally relaxed. The number of bands found on the gel gives the total number of super coils in a given DNA.
During replication, transcription, recombination, DNA repair and bending or folding of DNA or during gene expression or compacting of DNA as chromosomes, the normal relaxed DNA of 10.6 bp per turn, dramatically changes its base pairs per turn and the DNA is subjected to torsional stress; to accommodate such strain the DNA has to undergo super coiled states. Such topological changes in DNA can described by three parameters, they are
Linking number, Twist and Writhe:
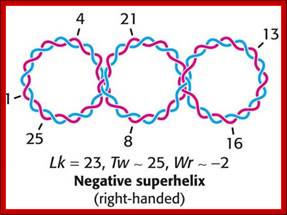
http://oregonstate.edu
Linking number (L):
It is an invariant but an integer value. It is the sum of two geometric components, one twist and the other writhe. Twist refers to the number of base pairs per one helical turn.
Twist (T):
The number of helical turns in a given DNA molecule of certain length, say in this case the DNA is double stranded, circular and 5300 bp. The number of twists = base pairs per turn or the helix. In a normal DNA, in this case, 10.6 bp/ turn, it amounts to 500 helical turns.
Writhe (W):
It is super coil (SC) that twists on its own length like a rubber band twist. The helical DNA crosses over on its own in space. The writhe can be plectonic or toroidal.
L = T + W;
When L < T + W, the DNA is in (-) SC state,
When L > T + W, the DNA is in (+) SC state,
L = T + W, where W = 0. Relaxed DNA;
Ex. DNA is ds, circular, 5300 bp long, bp/turn= 10.6, number of turns (360^0) = 500, Sc or writhe is Zero.
L = T + W;
Delta L = dT + dW,
Under wound= -dL = (-) SC
Over wound = +dL = (+) SC
Change in the state of DNA can be described by specific differences in linking numbers, i.e. D = dL / L (0),
So the L is; L = 500 + 0; so L = 500 turns, it also is also denoted as L (0)
If one wants to change the linking number, it is possible to do so, by nicking one of the strands, introduce certain number of turns, either way, then ligate the ends. The number of SC and handedness of the SC depends on the direction of turns and the number of turns. Let us take an example to explain this
Writhe and Negative super coils:
The DNA is 5300 bp,
The number of left handed turns (360^0) introduced is 50 that mean the number of turns removed is 50 (= 530 bp long DNA).
Calculate the number of writhes. In order to accommodate the loss of number of coils, it has to accommodate11.6 bp per turn, but energetically it is unfavorable, so it produces 50 negative super coils so as to have 10.6 bp/turn, as an unstrained molecule. The 50 negative super coils are 180^0 each and strands crossing over is on right handed direction.
T + W500 (5300
Writhe and Positive super coils:
The DNA is 5300 bp, 10.6bp/turn, so T=500 turns
The DNA is tuned right handed 50 times 360 ^0 each.
So the number of turns is 500+50=550, if this has to be accommodated the number of base pairs per turn will be=9.6; 5300%550 = 9.6, this bp is energy wise not compatible, so the DNA produces 50 positive super coils or writhes to be compatible. Each super coil is 180^0 and the DNA crossing is left-handed.
And L > L0
Generally, most of the times, positive super coils develop, when DNA opens and replication progresses in both directions; transcription also causes positive super coils. When DNA winds around histone protein complex in the left handed direction to form Nucleosomes, DNA experiences negative super coil state. Binding of proteins or protein complexes to DNA either during Replication or recombination or transcription, DNA is subjected to such stress and strains. Positive coils act as hindrance to the progress of replication fork as well as transcriptional fork and such positive super coils are removed and DNA is either rendered negative super coils or relaxed, which is essential for the said functions, otherwise DNA fails to replicate and fails to transcribe, which has serious consequences to the cell. In such situations DNA is subjected negative super coiling so the DNA easily opens ahead of the replication fork. Even ssDNA, such as circular DNA are in super coiled state, with out which the DNA fails to replicate and transcribe.
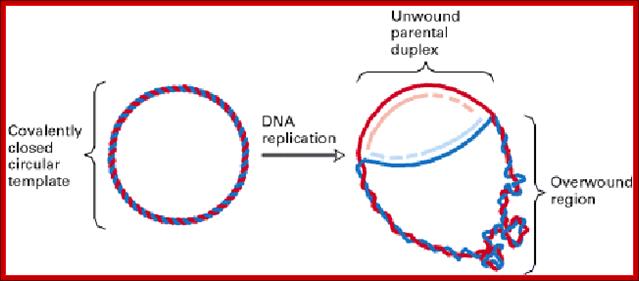
Opening of DNA causes over winding of DNA helix ahead of the replication fork, i.e positive super coiling; http://cienciasdejoseleg.blogspot.com/
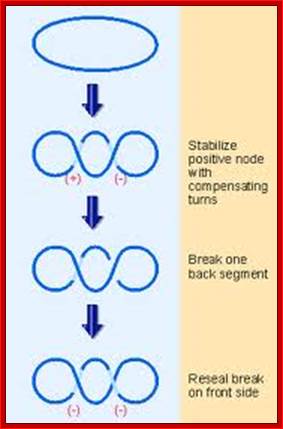
DNA gyrase may introduce negative supercoils in duplex DNA by inverting a positive supercoil. http://genes.atspace.org/
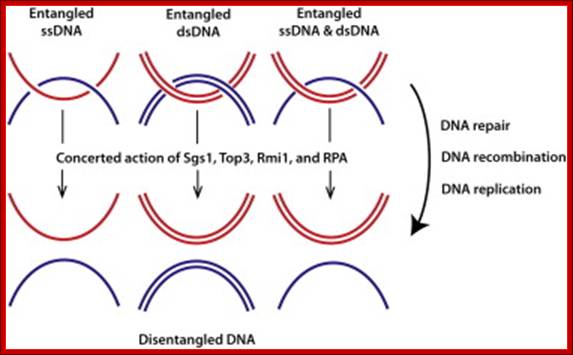
Catenation and decatenation; Petr Cejka et al; http://www.cell.com/
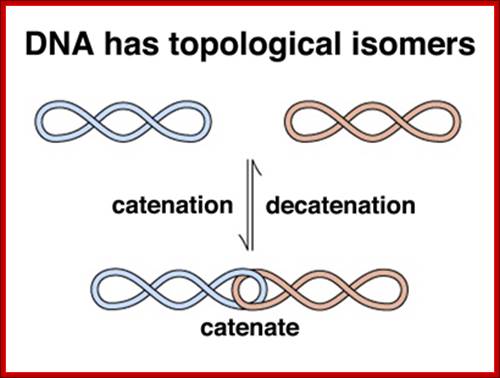
Biol 202 Genetics lecture10; http://www.discoveryandinnovation.com/
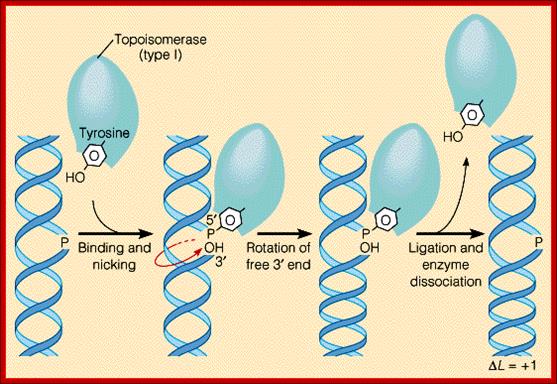
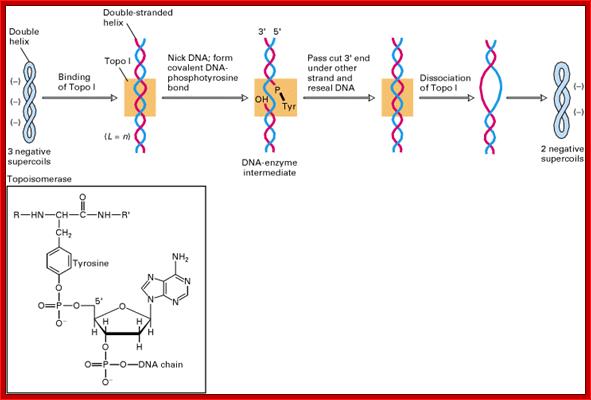
Topoisomerase I is a monomer with N and C ends 100kDa that bind to major groove and minor groove respectively. Then identifies and nicks it in such a way the 5’ end phosphate covalently links to its Tyrosine OH group. After relaxing the coil it covalently bonds the 5’phosphate end of the free OH group of the next nucleotide and covalently bonds. Topoisomerase I relaxing the Supercoiled,DNA: (www.Proteopedia.org); http://helicase.pbworks.com/
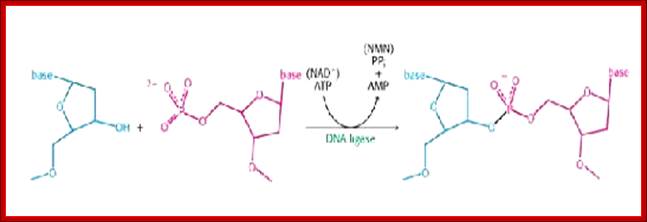
The Topoisomerase-1 has both nicking and ligase activity, here it shows ligase activity;
Action of Topoisomerase II;
Tyler-Huff; http://helicase.pbworks.com/
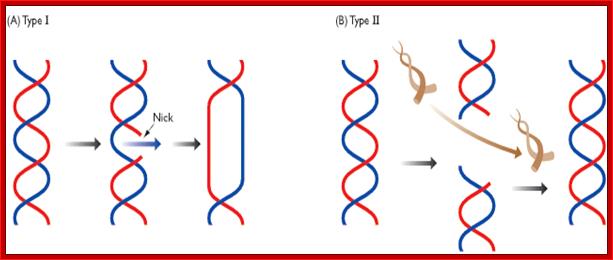
Action of Topo I cuts one of the two strands and Topo II cuts both the strands in staggered wy; http://biosiva.50webs.org/
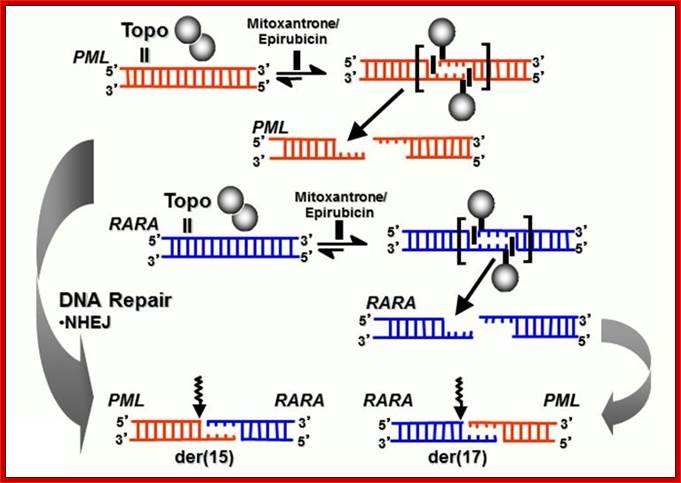
TopoII binds to specific region and cuts both strands in staggard way and recombine the strands; in the processes DNA is repaired; M. Joannides , et al; kings college London; http://www.mjhid.org/