Signal Transduction VI:
Signal Transduction in Plants:
Plants unlike animal systems are stationary but they react to environmental conditions and soil factors. Factors that affect or affect are light, temperature, moisture in the atmosphere, soil water, soil minerals, and toxic elements. Green plants respond to duration of the light in day-periodism; plants sense the day length and prevailing temperature they produce flowers, or undergo hibernation in their own ways and means. Plants as a large number of them are multicellular organized into different kinds of tissues, some are dead and some are living. All living cells, whatever kind and creed they belong, they do communicate with each other. Plants respond to light, for they have receptors and after absorption they transducer through various chemicals to initiate transcription for flowering. Growth hormones such as Auxins, though they are synthesized they are translocated and act on cells via auxin binding proteins, which rfegulate its downstream responses. Similarly GA which is synthesized in leaves is translocated and bring about changes either in flowering or seed/ grain germination. Ethylene is one gaseous substance generated within the plant and it has a tremendous effect on plant curvature, fruit ripening. Ethylene acts through its receptor sitting on plasma membrane. Similarly Abscissic acid responds to water conditions or drought and brings about changes in stomata closing or induce ripening and hasten falling of the leaves and flowers and fruits. In this chapter, only general picture of each of them are presented in the form of great diagrams from great authors, whom the author has acknowledged. Even soil bacteria like Agrobacterium or Rhizobia induce gall or nodule like structures by the nod factors binding to cell receptors and transducing signal for the development of nodules. Plant respond to pathogen infection in many ways and often use apoptosis as the last resort, so one can see burns in the leaves where pathogens are infected.
Plants also respond to viral and bacterial infection by the way of producing miRNA, dsRNA activated PKR and dsRNA activated RNaseL; all these leads to destruction of pathogens; in this course even plant cells die at the spot of infection; that is where one can see burnt spots in the leaves.

1) an overview of plant signal transduction, 2) sugar signaling, and 3) MAPK signaling in innate immunity. In addition, we are sharing our protocols, protoplast FAQ and virtual journal club. Genetic interactions between sugars and hormones, innate immune signalling activated by LRR receptors in Arabidopsis, www.molbio.mgh.harvard.edu
Light Induced Signal Transduction:
Light has tremendous effects on cellular activities including light triggered photosynthetic reactions, vision, photoperiodic induction of flowering, circadian rhythm and many others. In all these cells require light absorbing pigments such as chlorophyll/protein complex, phytochrome/protein complex, phototropin-protein complex; they inturn execute downstream reactions such as light harvesting and transducing the same to generate energy rich ATP and NADP molecules, activating genes for flower development and inducing photo-curvature (phototropism) movements respectively.
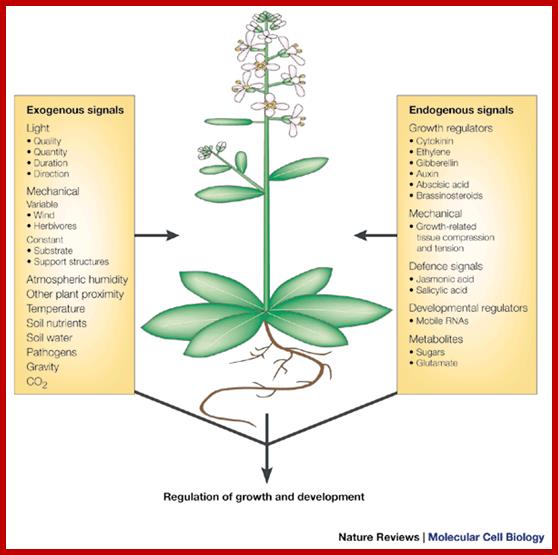
Signal processing and transduction in plant cells: the end of the beginning? The molecular elements of the plant sensory apparatus and signal-transduction systems can integrate these signals and reach a finely balanced decision as to how to grow and develop to most successfully survive and exploit the environment. As plant responses are generally irreversible growth responses, these signalling systems must compute each developmental decision with extreme care. Simon Gilroy and Anthony Trewavas; http://www.nature.com

Plants prepare to respond to light while still in the dark. Two proteins known as FHY3 and FAR1 bind to DNA in a plant cell and cause production of two other proteins known as FHY1 and FHL. When hit with far red light, a light-sensitive protein called phytochrome A (phyA) changes its shape. This shape change allows it to bind to FHY1 and FHL. FHY1 and FHL then carry the activated phyA into the cell nucleus. From there, phyA is able to initiate the plant's developmental responses to light such as growth, flowering and straining towards the light. Credit: Zina Deretsky, National Science Foundation; http://www.solutions-site.org/
How Plants Respond to Light


Godoy Herz MA, Kornblihtt AR, Barta A, Kalyna M, Petrillo E - Plant Signal Behav
Shedding light on the chloroplast as a remote control of nuclear gene expression. Plants use chloroplasts as light sensors that generate signals able to fine-tune nuclear gene expression. Light perceived in the chloroplast, the photosynthetically specialized organelle in the plant cell, triggers a signal that reaches the nucleus and affects alternative splicing, one important step in gene expression regulation able to generate several messages and proteins from a single gene. The light induced signal, or a derived one, is able to travel through the plant to non-photosynthetic tissue (i.e.: roots) affecting the alternative splicing there. https://openi.nlm.nih.gov

https://image.slidesharecdn.com/
https://image.slidesharecdn.com
The effects of light on plant growth and development:
Molecular genetic studies in Arabidopsis have identified four photoreceptor families that are present in all higher plants. There are three classes of blue light sensors: cryptochromes, phototropins and members of the Zeitlupe family. In addition the phytochromes enable plants to sense red and far-red light (Figure 1). In Arabidopsis these families are composed of two cryptochromes (cry1 and cry2), two phototropins (phot1 and phot2), three Zeitlupe-like sensors (ZTL, FKF1 and LKP2) and five phytochromes (phyA-phyE). Given that we mostly study phytochromes and phototropins-mediated signal transduction those two photoreceptors are presented in more detail.

A model for the regulation of shade avoidance by the phytochromes and PIF (Phytochrome Interacting Factor) bHLH-class transcription factors.Phytochromes are synthesized in their Pr conformation (PrB for phytochrome B in its Pr conformation). Upon light activation they are converted to their active Pfr conformation (PfrB for phytochrome B in its Pfr conformation), which accumulates in the nucleus. In sunlight characterized by a high R/FR ratio (see figure 3) the phytochromes are mostly in their Pfr conformation. Pfr specifically interacts with PIF transcription factors leading to their proteolytic degradation. In the shade, which leads to a reduction of the R/FR ratio, the phytochrome photo-equilibrium is pushed towards the Pr conformer. PrB no longer interacts with PIF transcription factors leading to their accumulation and transcription of shade marker genes.
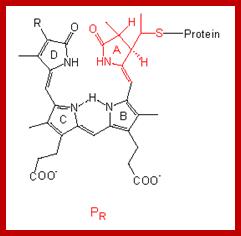

The structure of the linear tetrapyrrole is shown below. It is attached to the phytochrome protein through a sulfur linkage. http://www.mobot.org

https://webapps.molecular-networks.com
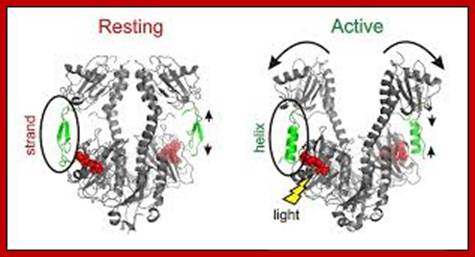
https://phys.org

ftp://godzilla.wnmu.edu/

essayon.pro/vernalization; m.blog.daum.net/kimuk

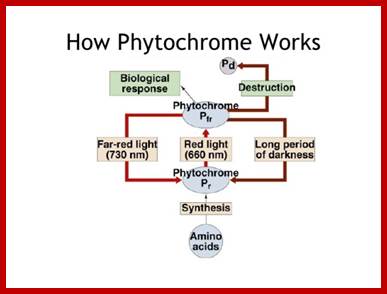
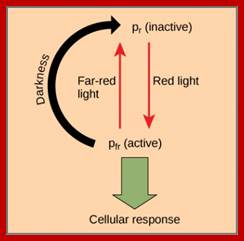
Phytochromephotoconversion; All phytochromes exist in two forms, called Pr and Pfr.a) Phytochromes are synthesised as Pr. Light absorption by Pr converts it to Pfr, initiating signals to the rest of the cell. Light absorption by Pfr converts it back toPr, b) Pr and Pfr have different absorption spectra. Pr absorbs light most strongly in the red region of the spectrum, whereas light absorption by Pfr is highest in the far red region. Therefore, plants exposed to red light contain mostly Pfr, while plants exposed to far red light contain mostly Pr. Phytochromephotoconversion; All phytochromes exist in two forms, called Pr and Pfr.a) Phytochromes are synthesised as Pr. Light absorption by Pr converts it to Pfr, initiating signals to the rest of the cell. Light absorption by Pfr converts it back to Pr.b) Pr and Pfr have different absorption spectra. Pr absorbs light most strongly in the red region of the spectrum, whereas light absorption by Pfr is highest in the far red region. Therefore, plants exposed to red light contain mostly Pfr, while plants exposed to far red light contain mostly Pr. http://intobiology.org.uk

http://www.bio.miami.edu

The effect of light on plant growth and development. All higher plants possess several classes of photoreceptors. Phytochromes (phyA-phyE) sense red and far-red light. Three distinct photoreceptor families: phototropins (phot1 & phot2), cryptochromes (cry1 & cry2) and the Zeitlupes (ZTL, FKF1 & LKP2) sense UVA/blue light. UVB-receptors are currently unknown. These photoreceptors allow plants to sense the intensity, quality, periodicity (day-length) and direction of light. These photoreceptors control important developmental transitions (e.g. the induction of flowering). Cryptochrome and phytochromes also determine whether a seedling will adopt an etiolated development (after germination in the dark) or a photomorphogenic development when the seedling develops in the light. The etiolated mode of development allows the seedling to rapidly emerge from the soil into the light. Shade avoidance and phototropism are two important adaptive responses, which allow seedlings to optimize photosynthetic light capture. The list of Arabidopsis photoreceptors is presented on this Fig. Christian Fankhauser; http://www.unil.ch/
Coupling photocatalysis and redox biocatalysis toward biocatalyzed artificial photosynthesis.
Chemistry - A European Journal 19:4392-4406 (2013) [IF=5.925]
Review paper highlighted as a concept article
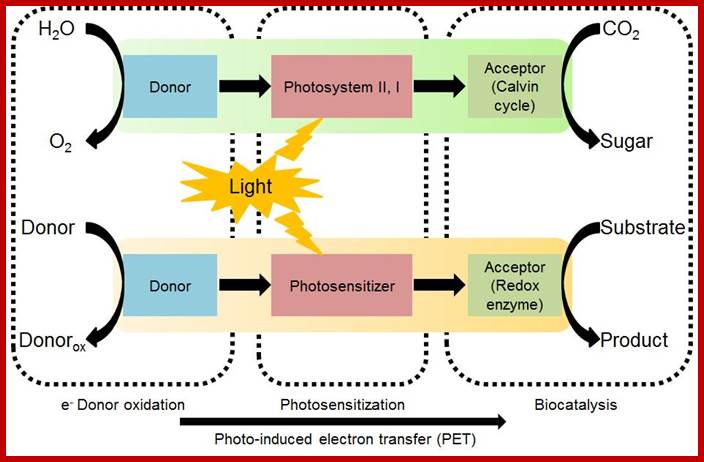
Solar energy utilization is accomplished in green plants through a cascade of photo-induced electron transfer, which remains a target model for realizing artificial photosynthesis. In this article, we introduce the concept of about how to design biocatalyzed artificial photosynthesis through coupling redox iocatalysis and photocatalysis to mimic natural photosynthesis. Key design principles for reaction components, such as electron donors, photosensitizers, and electron mediators, are described for artificial photosynthesis involving biocatalytic assemblies. Recent research outcomes that serve as a proof of the concept are summarized and current issues are discussed to provide a future perspective. S. H. Lee, J. H. Kim, and C. B. Park; http://biomaterials.kaist.ac.kr/
Floral initiation is orchestrated by systemic floral activators and inhibitors. This remote-control system may integrate environmental cues to modulate floral initiation. Recently, FLOWERING LOCUS T (FT) was found to be a florigen. However, the identity of systemic floral inhibitor or anti-florigen remains to be elucidated. Here we show that Arabidopsis thaliana CENTRORADIALIS (ATC), an Arabidopsis FT paralog, may act in a non-cell autonomous manner to inhibit floral initiation. Analysis of the ATC null mutant revealed that ATC is a short-day induced floral inhibitor. Cell type-specific expression showed that companion cells and apex expressing ATC are sufficient to inhibit floral initiation. Histochemical analysis showed the promoter activity of ATC mainly in vasculature but under the detection limit in apex, which suggests that ATC may moves from the vasculature to the apex to regulate flowering. Consistent with this notion, Arabidopsis seedling-grafting experiments demonstrated that ATC moved over long distance and that floral inhibition by ATC is graft transmissible. ATC probably antagonizes FT activity, because both ATC and FT interact with FD and regulate the same downstream meristem identity genes APETALA1, in an opposite manner. Thus, photoperiodic variations may trigger functionally opposite FT paralogs to systemically regulate floral initiation.

Systemic floral inhibition by a florigen paralog in Arabidopsis. Nien-Chen Huang, Wann-Neng Jane, Jychian Chen, and Tien-Shin Yu (2012); Plant Journal doi: 10.1111/j;

CONSTANS and the evolutionary origin of photoperiodic timing of flowering;
New model for the photoperiod response in plants. (A) The picture on the left represents the currently accepted model forArabidopsis, in which light-activated CO overcomes the temperature-dependent inhibition from FLC and induces the expression of FT in the phloem companion cells. FT is moved to the phloem and channelled to the apical meristem where it binds to FD and the complex is recruited into the nucleus. FT–FD binds to the promoter of SOC1 and other meristematic floral integrators, changing the vegetative developmental programme to the ABC programme, eventually producing flowers. (B) The model proposed here includes that depicted in A, but also recruits similar photoperiodic mechanisms to regulate other developmental programmes and basic physiological processes. Yellow arrows represent external signals: day/night transition; circadian clock; light quality; and a metabolic signal represented by a fertilizer bottle. Black arrows indicate some of the outputs of the photoperiodic response. http://jxb.oxfordjournals.org/
The functional phytochrome consists of two identical proteins, each with a chromophore. One part of the protein acts as the photoreceptor, and the other as akinase, which triggers cellular responses.

When the chromophore absorbs light, it isomerizes from one form to the other. This change in configuration results in a slight change in the kinase portion of the protein. http://www.bio.miami.edu/
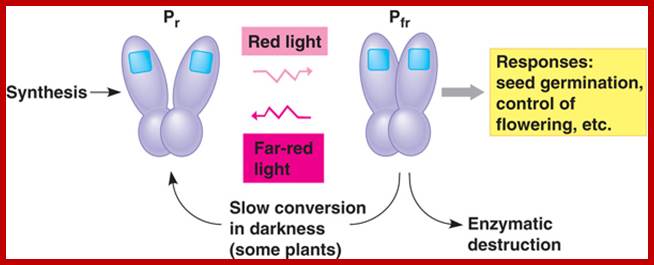
The kinase is the biologically active region of the molecule, and its interaction with other biological molecules elicits a physiological response. http://www.bio.miami.edu/


http://plantphys.info

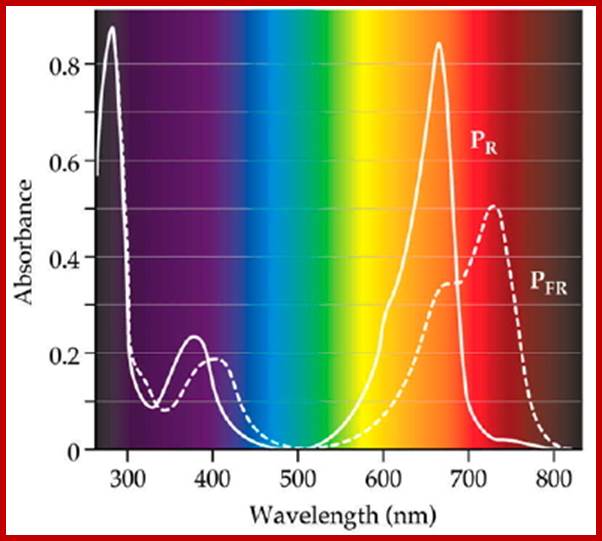
Diagram of interconversion of the Pr and Pfr forms of phytochrome. Obviously Pr absorbs red light 660nm very strongly, while pfr absorbs far-red light at 730nm, this leads to differences in their chemical structures and responses. http://www.photobiology.info/

You can see the relationship between chlorophyll and phytochrome, both have evolved from a tetrapyrrole ring system also seen in the phycobilin pigments of bacteria. The chromophore is bound to a protein, just as in the case of chlorophyll. The protein has a mass of 165 kilodaltons. What is interesting is how the chemical structure of phytochrome is altered to its complementary form when struck by photons of the correct energy level (wavelength!); http://plantphys.info/

It's all about gene expression. Ultimately, the signal transduction pathway results in the activation (or, in some cases, the suppression) of genes involved in the production of a specific, environmentally-induced phenotype.
A signal transduction pathway usually involves the increase in the activity of enzymes specific to a particular physiological response, either by stimulating transcription of the mRNA coding for that enzyme (transcriptional control) OR activating existing enzymes; http://www.bio.miami.edu/
Protein kinases drive adaptation responses:
Cellular signaling coordinates reprogramming of metabolism and gene expression and thus ultimately determines performance under stress conditions. Our work focuses on protein phosphorylation, which controls many cellular processes, especially those involved in intercellular communication and coordination of complex functions. We merge non-targeted and hypothesis-driven experiments and apply a broad range of genetic, metabolic, biochemical, physiological and genomic tools to gain novel insights into signaling mechanisms driving metabolic adaptation and gene expression in response to environmental stress. The GSK3/shaggy-like protein kinases MsK4 and ASKα are examples of protein kinases that we identified in a biochemical and a genetic screen as novel regulators of metabolic adjustment to high soil salinity. MsK4 associates with starch granules and positively regulates high salt tolerance by adjusting carbohydrate metabolism in response to environmental stress (Fig. 2).
ASKα links signal transduction with cellular redox balance during stress by regulating the glucose-6-phosphate dehydrogenase, a key enzyme for redox regulation (Fig. 3).
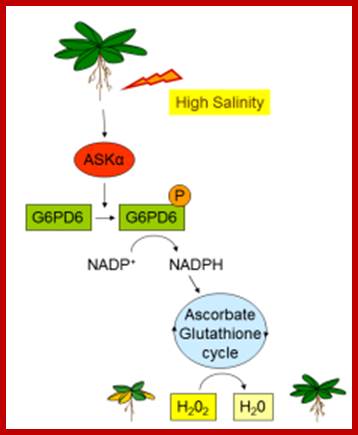
ASKα is an important regulator of ROS detoxification and, thus, acclimation to salt stress. High salinity activates ASKα, which, in turn, phosphorylates G6PD6, thereby stimulating its activity. Enhanced G6PD activity provides NADPH for the antioxidant system to remove excess ROS. Reduction of H2O2 to H2O can then be mediated by the glutathione peroxidase cycle or by the ascorbate-glutathione cycle. Claudia Jonak;http://www.gmi.oeaw.ac.at/
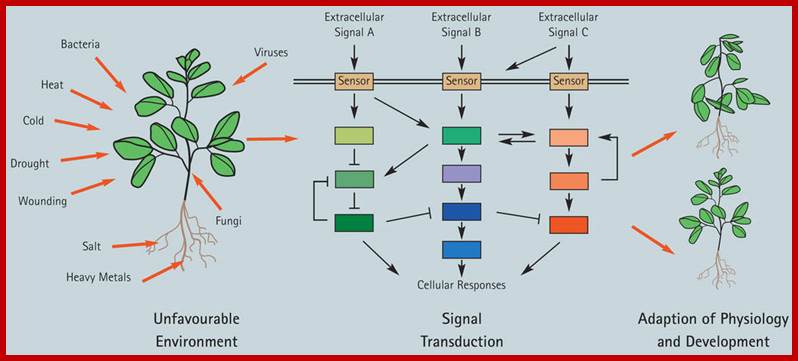
Plants respond to environmental stress. Plants are permanently exposed to a multitude of external stimuli; which plant cells have to transform into physiologically intelligible signals. Extracellular stimuli are perceived and internalized by various cellular receptors and are, subsequently, transduced by signaling cascades to induce appropriate cellular responses, which ultimately lead to physiological and developmental modifications determining the sensitivity or tolerance of a plant.; Claudia Jonak: http://www.gmi.oeaw.ac.at/
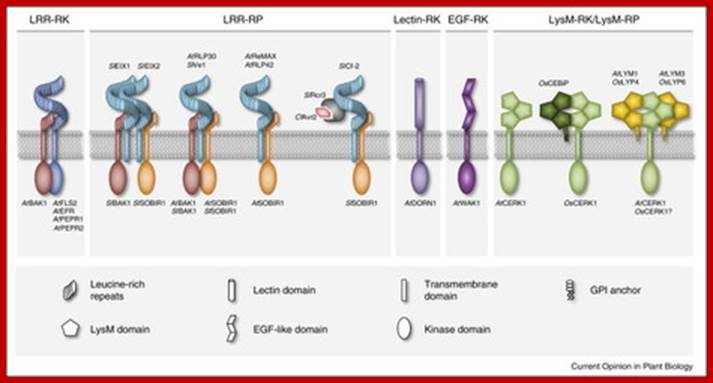
Immunity to microbial infection is a common feature of metazoans and plants. Plants employ plasma membrane and cytoplasmic receptor systems for sensing microbe-derived or host-derived patterns and effectors and to trigger inducible immune defenses. Different biochemical types of plasma membrane immune receptors mediate recognition predominantly of peptide and carbohydrate patterns. Current research highlights the role of immune receptor complex formation in plant immunity. In particular, ligand binding by immune receptors generates molecular surfaces that enable either receptor homo-dimerization or co-receptor recruitment for subsequent signal transduction. New insight into negative regulatory principles of immune receptor function further suggests substantial dynamics in protein–protein interactions at the plasma membrane that we are only beginning to understand. From www.sciencedirect.com - May 21, 5:43 AM; http://www.scoop.it/
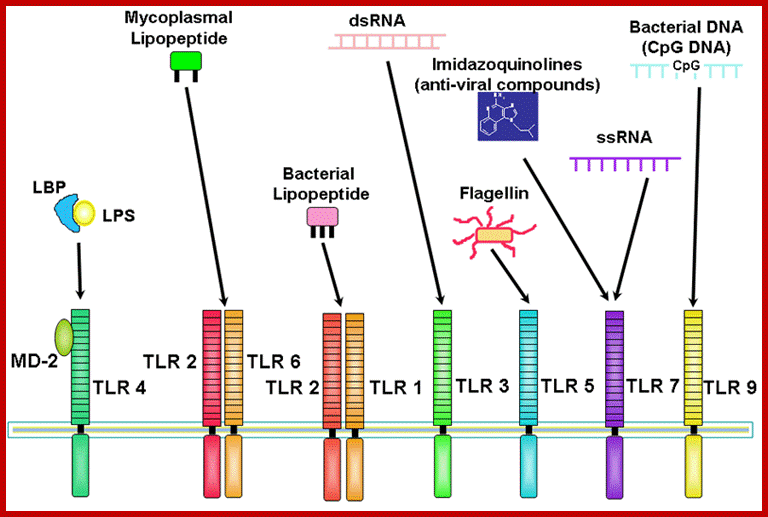
Plants are equipped with receptors for different kinds of pathogens. http://biotec.campusnet.unito.it/
Outline of Research and Education; Plant immunity against?
In nature, plants cope with a wide range of microbes that reside on the surface of or within the plant tissues. Plants disregard or tolerate the presence of these plant-inhabiting microbes at non-damaging levels, despite an elaborate innate immune system to detect and repel microbes. We hypothesize that plant immunity senses and reacts to “danger” signals (DAMPs) generated upon pathogen challenges over the background microbial signals (MAMPs). We aim to decipher the molecular principles and mechanisms underlying this sophisticated function of plant immunity, with major focuses on immune sensors and signaling, defense-related transcriptional reprogramming, and microbial strategies to avoid/dampen host immune activation. We believe that our studies will gain important insight into general principles of plant immunity, and thus offer new effective approaches for controlling plant health and growth in sustainable agriculture

Plant immunity: Pattern recognition receptors and co-receptors in plants; Yusuke SAIJO; http://bsw3.naist.jp/
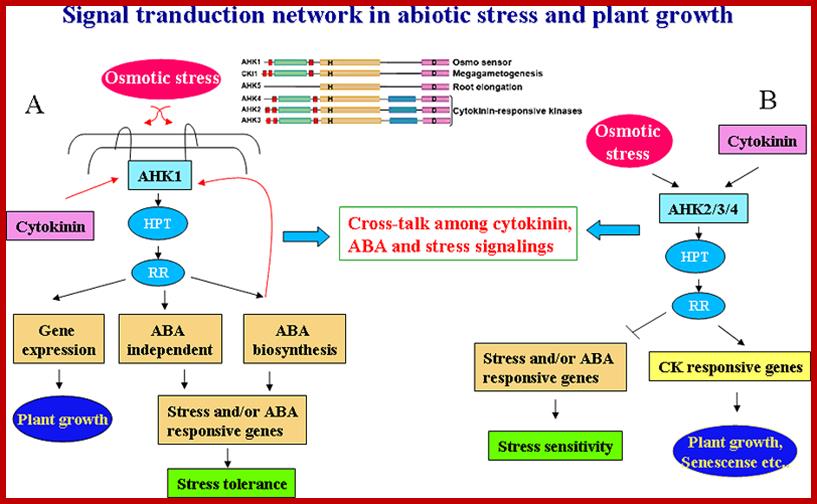
A generalized diagram shows interaction of plants with biomolecules in response to stresses

New checkpoints in stomatal defense.
Trends Plant Sci. 2013 Apr 10;
· An oxylipin-dependent signaling pathway controls stomatal defense.
· Abscisic acid modulates the biotic stress signal in guard cells.
· Distinct mitogen-activated protein kinase modules regulate biotic and abiotic stresses.
· Distinct NADPH oxidase isoforms control reactive oxygen species in response to biotic and abiotic stresses.
- Salicylic acid is a key signaling step required for stomatal defense.; http://www.versailles.inra.fr/
Recent reports have revealed new guard


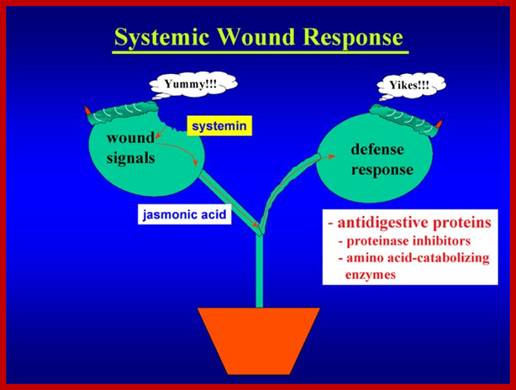
Signal perception and signal transduction through MAPK pathways; Wound response in tomato plant, a paradigm for plant responses; http://ww2.biol.sc.edu/
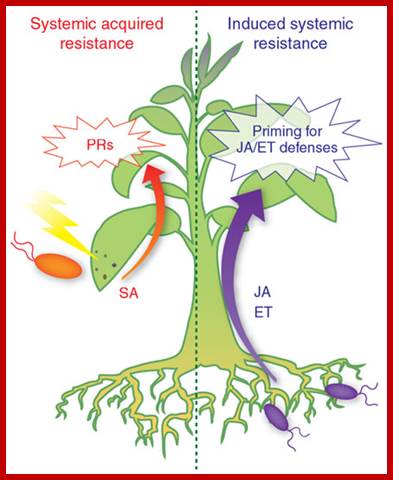
Networking by small-molecule hormones in plant immunity; Plants live in complex environments in which they intimately interact with a broad range of microbial pathogens with different lifestyles and infection strategies. The evolutionary arms race between plants and their attackers provided plants with a highly sophisticated defense system that, like the animal innate immune system, recognizes pathogen molecules and responds by activating specific defenses that are directed against the invader. Recent advances in plant immunity research have provided exciting new insights into the underlying defense signaling network. Diverse small-molecule hormones play pivotal roles in the regulation of this network. Their signaling pathways cross-communicate in an antagonistic or synergistic manner, providing the plant with a powerful capacity to finely regulate its immune response. Pathogens, on the other hand, can manipulate the plant's defense signaling network for their own benefit by affecting phytohormone homeostasis to antagonize the host immune response. Corné M J Pieterse1, Antonio Leon-Reyes1, Sjoerd Van der Ent1 & Saskia C M Van Wees; http://www.nature.com/
Phytochrome induced G-response:
Phytochrome bound protein complex absorb Red light wave length and transforms into auto phosphorylated Phy-Pfr. This is an active complex which transduces its signal pathway into the nucleus, where it binds to specific receptors such as HFR-PF3 and activates genes like Hy5, CCA1 and LHY.
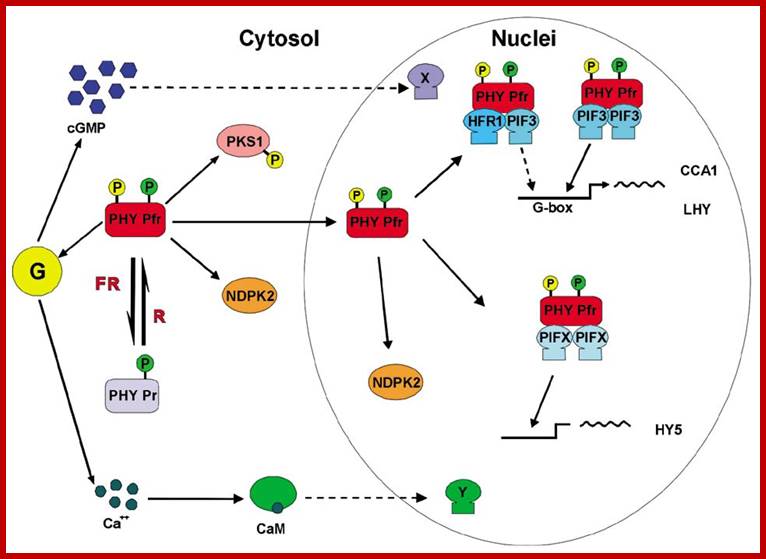
Physiology of flowering; Plant Cell Biol; http://b.21-bal.com
The same induces flowering by what is called flower inducing responses. The activated Phy-Pfr also activates G-protein leading to the production cGMP. It also activates of Ca2+ channels.
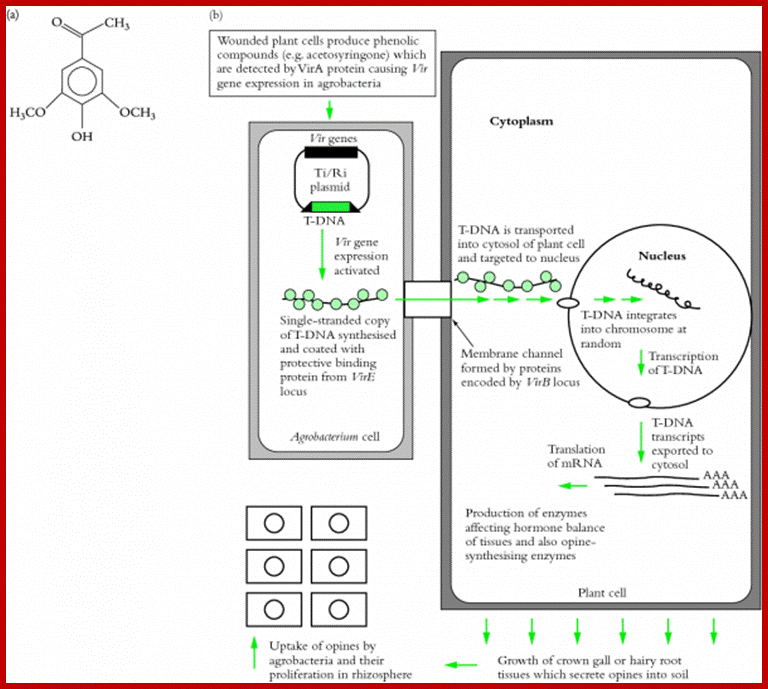
Agrobacterium tumefacians is a plant gall inducing bacteria. It binds to plant cell wall by receptor ligand contact and gets activated, by opines produced by plants. This is where many bacterial T-DNA genes express in a cascade to generate proteins that are responsible for cutting one strand of the DNA at the right end of T-DNA and peel of the strand and coats with proteins and the same is transported through a channel created by its gene products that connects bacterial cell wall and host cell wall. This T-DNA gets entry into the nucleus, where it gets integrated and expresses products responsible for crown gall formation.
Infection of Plants with pathogens and plant responses:
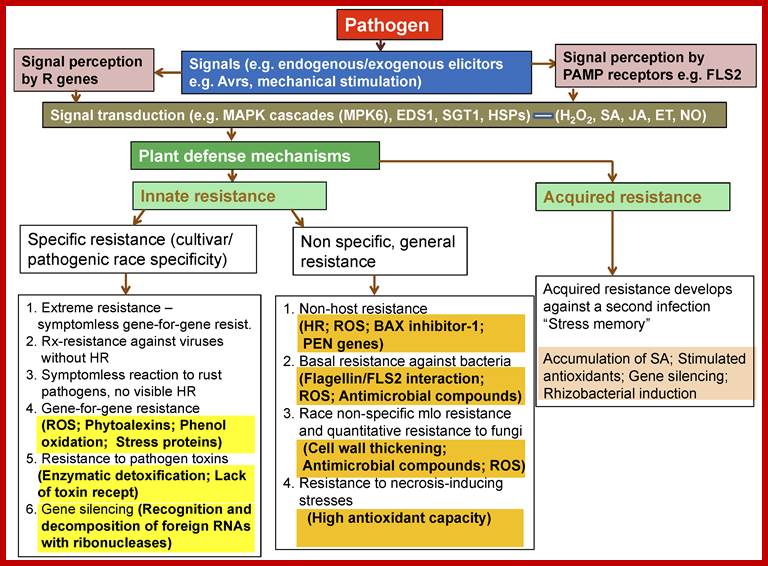
Plant Tolerance to Biotic Stresses; By Geoffrey Onaga and Kerstin Wydra; https://www.intechopen.com/
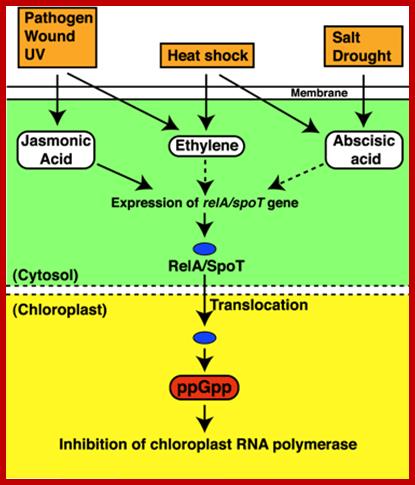
In response to stress(s) like pathogen infection, heat shock, salt stress or drought induce the synthesis of jasmonic acid, ethylene and ABA respectively. They in turn activate genes such as relA/spoT genes. These products enter into chloroplasts where they produce ppGpp and inhibit chloroplast RNA polymerase. This action is similar to that of bacterial Rel A protein action. Identification of the bacterial alarmone guanosine 5′-diphosphate 3′-diphosphate (ppGpp) in plants; http://www.pnas.org
In response to stress(s) like pathogen infection, heat shock, salt stress or drought induce the synthesis of jasmonic acid, ethylene and ABA respectively. They in turn activate genes such as relA/spoT genes. These products enter into chloroplasts where they produce ppGpp and inhibit chloroplast RNA polymerase. This action is similar to that of bacterial Rel A protein action.
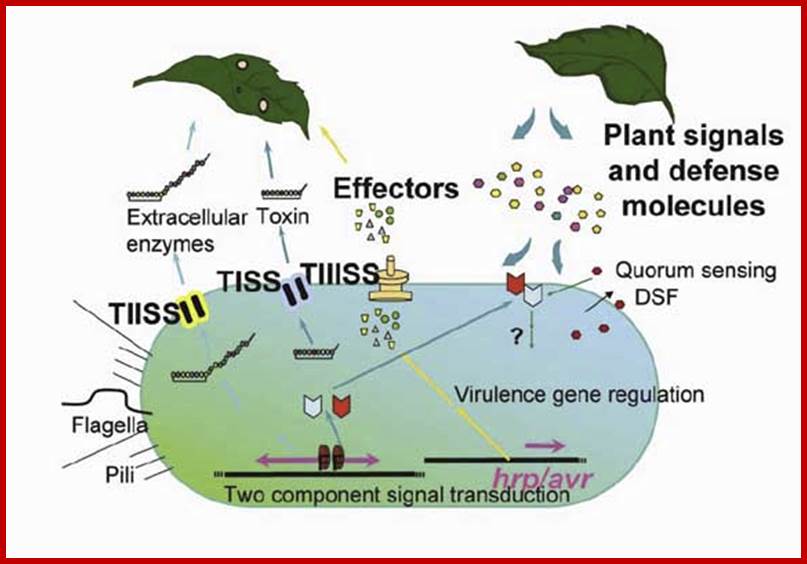
http://www.enseignegambetta.com/
Auxins:
Auxins are the largest group of phytohormones. Auxins aere the first to be discovered. Auxin enters extracellular space of te plant cell wall and binds to Auxin receptor, which transports the same into cytoplasm. This leads to the activation of specific genes in response to auxins.

Auxin in action: signalling, transport and the control of plant growth and development. http://www.researchgate.net
Hormones have been at the centre of plant physiology research for more than a century. Research into plant hormones (phytohormones) has at times been considered as a rather vague subject, but the systematic application of genetic and molecular techniques has led to key insights that have revitalized the field. In this review, we will focus on the plant hormone auxin and its action. We will highlight recent mutagenesis and molecular studies, which have delineated the pathways of auxin transport, perception and signal transduction, and which together define the roles of auxin in controlling growth and patterning. ; http://www.nature.com/
Schematic representation of NO–cytokinin antagonistic interactions.

(1) Certain cytokinin species such as zeatin may chemically react with peroxynitrite (ONOO-), producing derivates with virtually no biological activity. (2) NO might also negatively impact cytokinin signaling since the protein HISTIDINE PHOSPHOTRANSFER PROTEIN 1 (AHP1), a key element in the phosphorelay mechanism involved in cytokinin transduction inArabidopsis, may undergo S-nitrosylation at cys-115, rendering this protein incapable of transferring phosphoryl groups from the cytokinin receptors to the ARABIDOPSIS RESPONSE REGULATORs (ARRs). Protein S-nitrosylation and phosphorylation are represented by “–S–NO” and “P,” respectively. http://journal.frontiersin.org/
http://www.frontiersin.org/
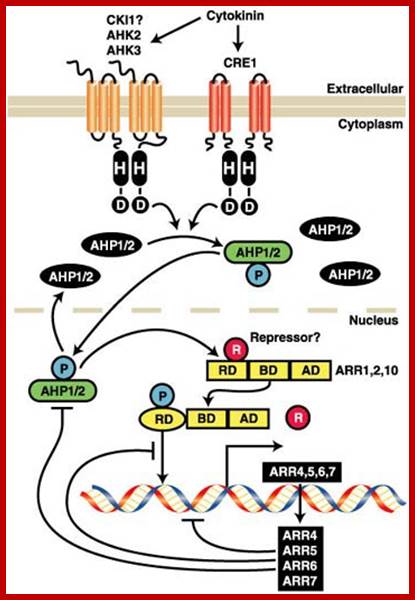
Cytokinin Signaling Pathway: Cytokinins bind to the cell surface receptors which get activated and the receptor has histidine kinase domain. This ultimately activates specific genes. Activation leads to the expression of set of genes; http://molbio.mgh.harvard.edu/
Ethylene

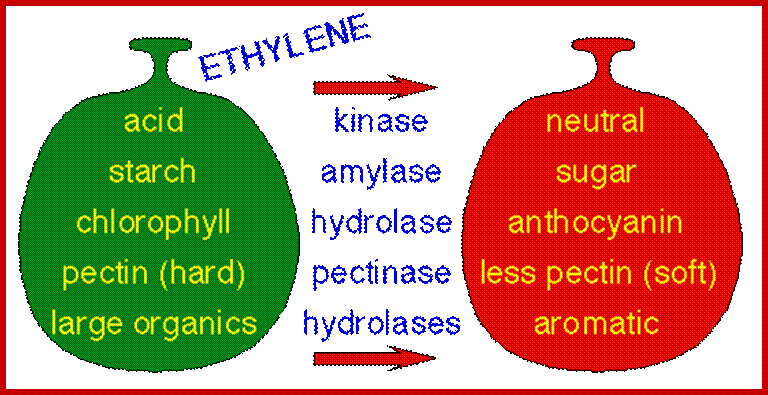

With the exception of ethylene, the signaling pathways that regulate fruit ripening remain largely undefined; http://gcat.davidson.edu/

Fruit ripening induced by ethylene. http://www.ext.colostate.edu/
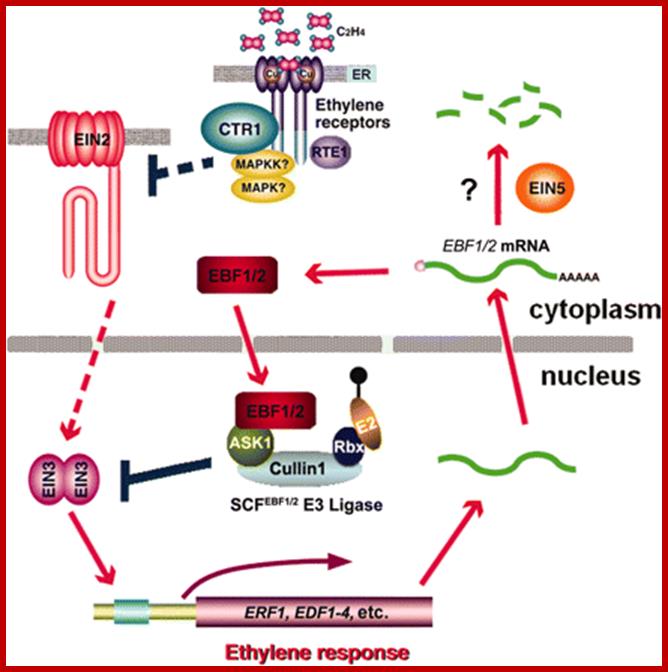
Ethylene response cascade; Ethylene enters cells easily and binds to its receptors. The receptors act upon specific genes and express the same. http://ucce.ucdavis.edu/
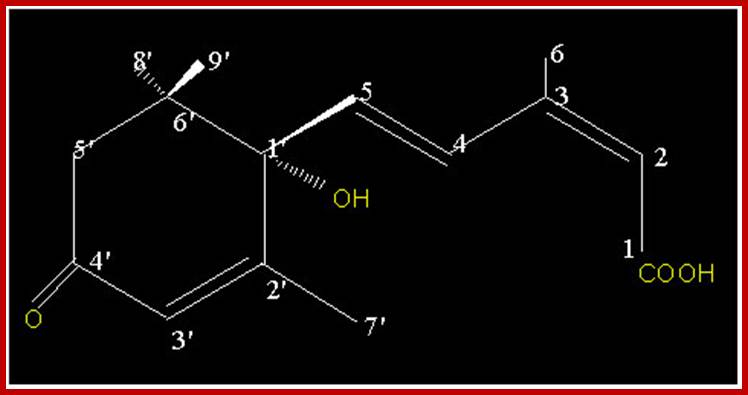
ABA
Water circulation:
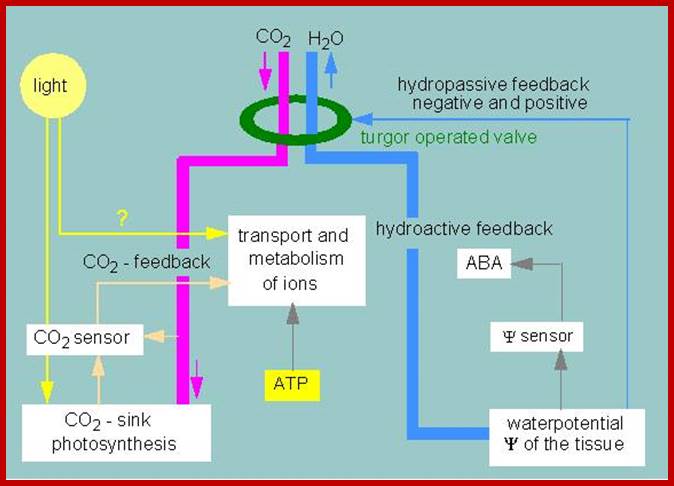
http://kfrserver.natur.cuni.cz/
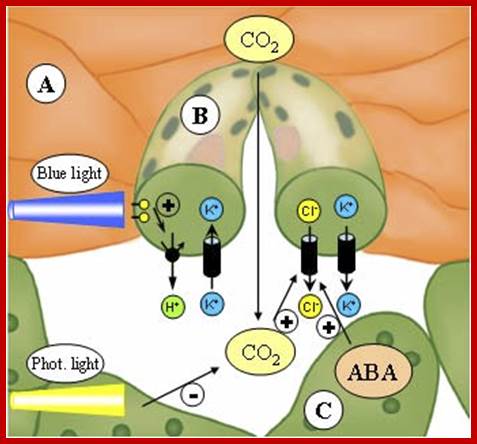
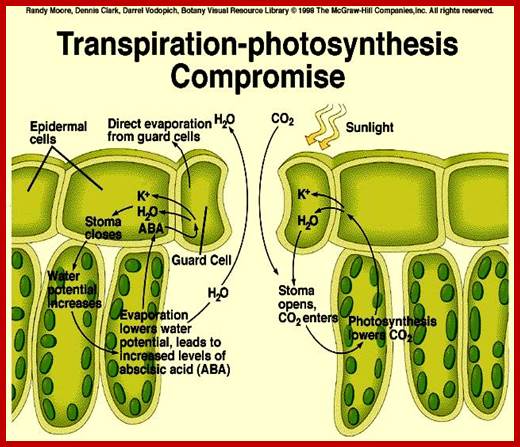
The opening and closing of the stomata allows the sending of signals to the palisade layer so that it can decide how much food is needed.; This is done when stomata allows for the flow of air into the spongy mesophyll which then sends signals to the palisade layer. Guard cells also control the air exchange of CO2 and O2. When photosynthesis is high more O2 is used and the byproduct CO2 is release. Guard cells control the gaseous exchange between CO2 and O2 during photosynthesis in the palisade layer; Guard cells send signals; http://naribiochemwiz007.wordpress.com/
Light induced opening and closing of stomata regulated by the activation ion-gated channels such as K+ ions in exchange with H+ ions (leading to opening) and ABA induced expulsion of K/Cl ions in closing the stomata.
Factors that speed up transpiration will also increase the rate of water uptake from the soil. If the loss of water is faster than the rate at which it is being replaced by the roots, then plants can slow down the transpiration rate by closing some of their stomata. This is regulated by guard cells, which lie on either side of a stoma.
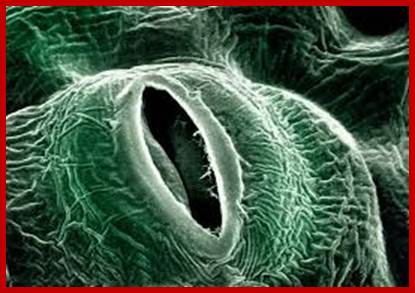
The science of the stomata of plants; https://plantstomata.wordpress.com
A doorman in plant cells:
The Research Group of Klaus Harter at the Centre for Plant Molecular Biology (ZMBP), University of Tubingen, Germany, identified the AHK5 as an important signalling Protein for the stress-response in plant cells. The results were published in PLoS ONE on 18 June 2008.
Stomatal guard cells monitor and respond to environmental and endogenous signals such that the stomatal aperture is continually optimized for water use efficiency. A key signalling molecule produced in guard cells in response to plant hormones, light, carbon dioxide and pathogen-derived signals is hydrogen peroxide (H2O2). The mechanism by which H2O2 integrates multiple signals via specific signalling pathways leading to stomatal closure is not known.
Stomatal apertures on the upper side of plant leaves enable gas exchange (Photo: Uni Basel, WBI Freiburg)
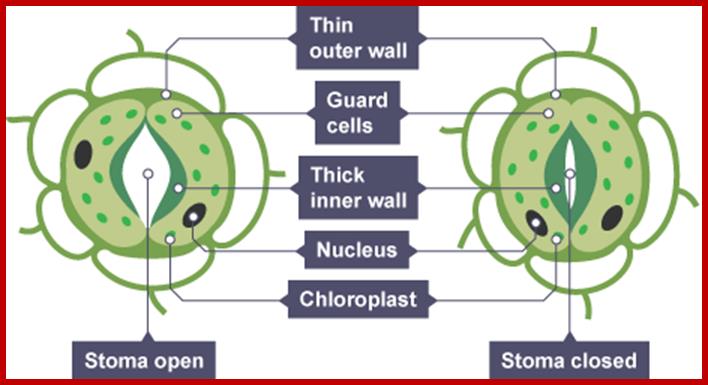
http://www.bbc.co.uk/
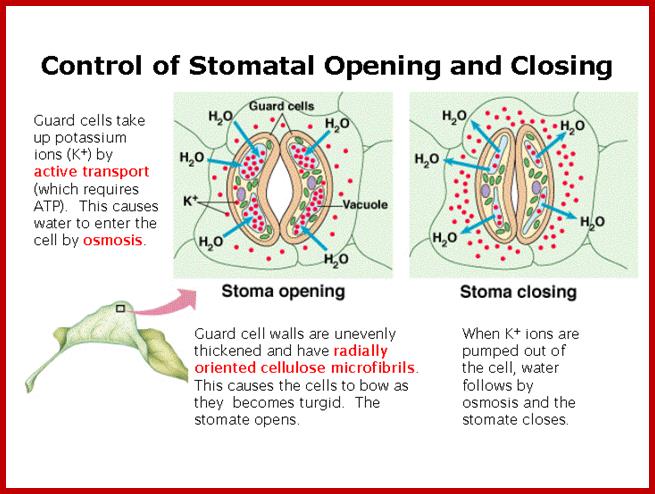
If the guard cells are turgid, then they curve forming ‘sausage-shaped’ structures with a hole between them. This is the stoma.
However, if the guard cells are flaccid due to water loss, they shrivel up and come closer together, closing the stoma. This is turn reduces the water loss due to transpiration, and can prevent the plant from wilting; ;http://www.bbc.co.uk/
Preuniversity.com; http://www.pinsdaddy.com

Points of resistance to the diffusion of CO2 from outside the leaf to the chloroplasts (source: Taiz L., Zeiger E., 2010);http://www.tankonyvtar.hu/
http://usdbiology.com/

GC membrane –transport process during opening left and closing right. http;//www.bio.fsu.edu
In essence stomatal openings initiated by H+ extrusion, which hyperpolarizes the PM and acidifies the apoplast. The effects increase the driving force for uptake of K+ions and activate voltage regulated K+ in channels. Cl- and suc may also be taken up. H+ pumping into vacuoles provides for K+ antiport and Cl uptake. GC solute increase leads to H2O uptake and therefore an increase in GC volume and aperture size, stomal clsing is initiated by activation of A- channel, which depolarizes he PM. This effect increases for K+ efflux and activates the volatage –regulated K+ out channel. Several channels provide for solute release from the vacuole. Internal and external ABA receptors cause the elease of Ca2+ though cellula messengers, eg IP3; ABA also causes Ca2+ influx. ABA also induces stotal closure by Ca2+ independent means. Compartments are not to the scale.

Guard cell abscisic acid signalling and engineering drought hardiness in plants;
a, ABA is detected by as yet unidentified receptors (right guard cell) and induces cytosolic Ca2+ elevations (1) through extracellular Ca2+influx and release from intracellular stores (for reviews see refs 2, 8, 9). (2) [Ca2+]cyt elevations activate two types of anion channel that mediate anion release from guard cells, slow-activating sustained (S-type) or rapid transient (R-type) anion channels3, 8. (3) Anion efflux causes depolarization, which activates outward-rectifying K+ (K+out) channels and results in K+ efflux from guard cells2, 3. (4) ABA causes an alkalization of the guard cell cytosol, which enhances K+out channel activity44. Overall, the long-term efflux of both anions and K+ from guard cells contributes to the loss of guard cell turgor, leading to stomatal closing3. Over 90% of the ions released from the cell during stomatal closing must be first released from vacuoles into the cytosol. (5) At the vacuole, [Ca2+]cyt elevation activates vacuolar K+ (VK) channels, which are thought to mediate Ca2+-induced K+ release from the vacuole8. In addition, fast vacuolar (FV) channels can mediate K+ efflux from guard cell vacuoles at resting [Ca2+]cyt45. ABA also inhibits ion uptake, which is required for stomatal opening (left guard cell). (6) [Ca2+]cyt elevations inhibit the electrogenic plasma membrane proton-extruding H+-ATPases46 and K+ uptake (K+in) channels2, 3, 44 (7). Initiation of ion efflux (1–5, right guard cell) and inhibition of stomatal opening processes (6, 7, left guard cell) provide a mechanistic basis for ABA-induced stomatal closing8.
b, Ca2+-releasing second messengers that mediate ABA-induced stomatal closing. Second messengers have been implicated in ABA signalling in guard cells, including InsP3 and cADPR. (8) Biochemical evidence indicates that ABA stimulates InsP3 production in guard cells2. (9) InsP3 can release Ca2+ from intracellular stores, inhibit K+in channels and cause stomatal closure9, 13, 22, 44. (10) [Ca2+]cyt elevations may be amplified by a Ca2+-induced Ca2+ release (CICR) mechanism from the vacuole through activation of the Ca2+-dependent slow vacuolar (SV) channel8, 45. Data questioning this SV model for CICR47 and recent other data supporting it48 indicate the potential for molecular genetic analyses. (11) ABA stimulates the production of cADPR in plant cells49, which also increases [Ca2+]cyt in guard cells20, 45 and promotes stomatal closing. Julian I. Schroeder, June M. Kwak and Gethyn J. Allen
Retrograde signaling pathways and chloroplast–mitochondrion cross-talk in higher plant cells: Coordination of gene expression between organellar and nuclear genomes: Jesse D. Woodson & Joanne Chory.

Retrograde signalling pathways and chloroplast–mitochondrion cross-talk in higher plant cells. Jesse D. Woodson & Joanne Chory; http://www.nature.com/
This figure depicts chloroplast-to-nucleus and mitochondrion-to-nucleus retrograde signaling pathways in the higher plant cells (and in yeast and animals where noted). Seven different pathways are highlighted. a | The use of chloroplast-generated reactive oxygen species (ROS) to induce nuclear gene transcription. b | Control of nuclear gene regulation by the redox state of the photosynthetic electron transport chain (PET). c | Chloroplast Mg–protoporphyrin IX (Mg–proto) accumulation. d | Inhibition of plastid gene expression (PGE). c and d lead to the repression of nuclear-encoded chloroplast protein genes. Signals from inhibited mitochondrial gene expression act synergistically with the PGE pathway. Two putative Mg–proto signaling pathways are depicted: in pathway 1, GENOMES UNCOUPLED 1 (GUN1) or a putative GUN1-dependent chloroplast protein (GDCP) facilitate the export of Mg–proto from the chloroplast where it interacts with cytoplasmic signalling factors; in pathway 2, GUN1 or GDCP sense Mg–proto accumulation and other retrograde signals within the chloroplast and send an unidentified signal to the nucleus to control transcription of chloroplast protein-encoding genes. e | Mitochondrial electron transport chain (mtETC) dysfunction leads to transcriptional changes in the nucleus in several phyla. f | An aberrant mitochondrial protein leads to cytoplasmic male sterility (CMS) by affecting nuclear gene expression. g | Mitochondrial haem synthesis as a cellular sensor for O2 availability in yeast. Proteins that are known to be involved in these pathways are designated as ovals. ABI4, abscissic acid insensitive 4; AOX, gene encoding the mitochondrial alternative oxidase; HAP1, haem activation protein; LHCB, gene encoding photosystem II chlorophyll a/b-binding protein; PQ, plastoquinone; PSI, photosystem I; ROX1, repressor of hypoxic genes; STN7, a thylakoid protein kinase.
Analysis of calcium signaling pathway in Plants☆ Oliver Batistič,;Jörg Kudla
Calcium serves as a versatile messenger in many adaptation and developmental processes in Plants Ca2 + signals are represented by stimulus-specific spatially and temporally defined Ca2 + signatures. These Ca2 + signatures are detected, decoded and transmitted to downstream responses by a complex toolkit of Ca2 + binding proteins that function as Ca2 + sensors.
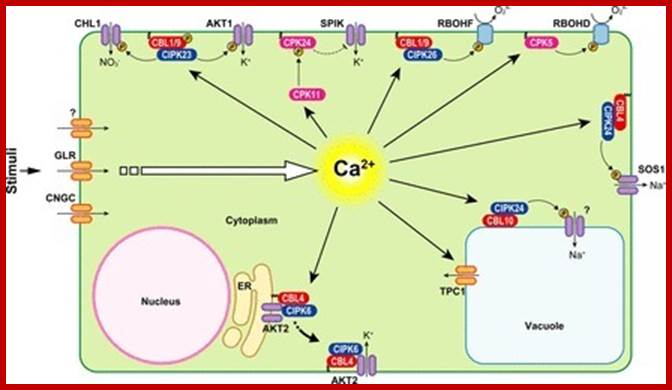
Ca2 +-mediated signal transduction processes in plant cells. Examples of signal transduction processes mediated by CaMs, CDPKs and CBL–CIPK complexes together with their respective target proteins, which were investigated in different plant species (indicated by the acronyms: At, Arabidopsis; St, potato; Nt, tobacco). See text for further details. Phosphorylation of target proteins is indicated by the yellow circled P. http://www.scoop.it/
The AHK5 protein detects stress; Prof. Dr. Klaus Harter:
Although
AHK5 is expressed at low levels in guard cells, we identify a unique role for
AHK5 in stomatal signalling. Arabidopsis mutants lacking AHK5 show reduced
stomatal closure in response to H2O2, which is reversed by complementation with
the wild type gene.
Over-expression of AHK5 results in constitutively less stomatal closure.
Abiotic stimuli that generate endogenous H2O2, such as darkness, nitric oxide
and the phytohormone ethylene, also show reduced stomatal closure in the ahk5
mutants. However, ABA caused closure, dark adaptation induced H2O2 production
and H2O2 induced NO synthesis in mutants. Treatment with the bacterial pathogen
associated molecular pattern (PAMP) flagillin, but not elf peptide, also
exhibited reduced stomatal closure and H2O2 generation in ahk5 mutants.
These findings identify an integral signalling function for AHK5 that acts to
integrate multiple signals via H2O2 homeostasis and is independent of ABA
signalling in guard cells.
Tyrosine phosphorylation in plant cell signaling; Sheng Luan*
Tyrosine Phosphorylation in Plants; http://www.pnas.org/
A simplified model of ABA-induced stomatal closure (see more detail in refs. 19 and 20). ABA receptors are possibly located in the plasma membrane and/or the cytoplasm. ABA signal is transmitted through calcium-dependent and calcium-independent pathways, leading to the regulation of ion channel activities in the plasma membrane (PM) and Tonoplast. A tyrosine phosphatase (PTP) appears to be located downstream of calcium and regulates tonoplast K-channels responsible for K efflux from the vacuole. Vacuole K efflux and PM channels contribute to turgor decrease, leading to stomata closure.
Model for signalling processes in the maintenance and/or the differentiation of procambial cells and xylem cell precursors:
Signals that control plant vascular cell differentiation;

Model for signalling processes in the maintenance and/or the differentiation of procambial cells and xylem cell precursors. ;Hiroo Fukuda; http://www.nature.com
In procambial cells (PCs), the coordinated signalling by cytokinin and auxin induce the expression of genes that are involved in the maintenance of procambial activities. The auxin-signalling pathway might involve gene expression of auxin-response factors, such as MONOPTEROS (MP), that also function as transcriptional activators, and their repressors, the AUX/IAA proteins. Cytokinin might be perceived by the WOL/CRE1/AtHK4 cytokinin receptor, which, in turn, transmits an intracellular signal that is mediated by a His–Asp phosphorelay mechanism to PC-related histidine-containing phosphotransfer factors (AHPs) and then to PC-related type-B response regulators (ARRs). The type-B ARRs might function as transcriptional activators of PC-related genes including the genes of their repressors, the type-A ARRs. The presence of repressors in auxin- and cytokinin-signalling pathways might allow cytokinin and auxin signalling to be temporal. Brassinosteroids (BRs in the figure) are biosynthesized actively in PCs and secreted, but brassinosteroids do not work as a signal for the maintenance of procambial activities. Instead, brassinosteroids, in the presence of auxin, might initiate differentiation of procambial cells to precursors of xylem cells (pXCs) after recognition by a receptor, which might be a heterodimer, composed of either brassinosteroid-insensitive-1 (BRI1) or one of the BRI1-like proteins (BRL1–BRL3), plus BRI1-associated receptor kinase-1 (BAK1). The brassinosteroid signal inactivates the negative regulator BIN2 (brassinosteroid-insensitive-2), which allows the unphosphorylated form of bri1-EMS-suppressor-1 (BES1) and brassinazole-resistant-1 (BZR1) to translocate to the nucleus and to promote pXC-related gene expression. Among the most important pXC-related genes that are induced by brassinosteroids might be the HD-ZIP-III-homeobox gene family, which might function in further xylem cell differentiation. KANADI and the microRNAs MIR165 and MIR166 might suppress differentiation of PCs to pXCs. The suppression by the microRNAs might be caused by the rapid degradation of the HD-ZIP-III gene mRNA through RNAi machinery.
Herbivore-induced, indirect plant defences: Gen-ichiro Arimura, Christian Kost, Wilhelm Boland

https://www.researchgate.net
Schematic representation of the signalling pathways required for herbivore-induced responses in plants. This scheme merges the evidence obtained from several plant taxa. The overall scenario may differ in certain plants; in particular the existence and the extent of synergistic and antagonist interaction between pathways may vary significantly. Elements in blue represent enzymes. Broken arrows indicate possible steps not yet described. Abbreviations: ACC, 1-aminocyclopropane-1-carboxylic acid; ACS, ACC synthase; DAG; diacylglycerol; FAC, fatty acid-amino acid conjugate; FAD, ω-3 fatty acid desaturase; HIPV, herbivore-induced plant volatiles; JA, jasmonic acid; JMT, JA carboxyl methyl transferase; LOX, lipoxygenase; MAPK, mitogen-activated protein kinase; MeJA, methyl JA; MeSA, methyl SA; OPDA, 12-oxophytodienoic acid; PL, phospholipase; PA, phosphatidic acid; SA, salicylic acid; SAM, S-adenosyl-methionine; SAMT, SA carboxyl methyl transferase; TF, transcription factor.
Stomatal Responses to DroughtStress and Air Humidity: L. E. Arve1, S. Torre1, J.E. Olsen1 and K. K. Tanino2
Stomatal signaling and movement:
Stomatal closure occur when the two guard cells surrounding the stomatal opening lose turgor pressure and close the opening (Outlaw, 2003). There are many signals that induce stomatal closure, among these the best known signal is probably ABA. In the signaling pathway towards stomatal closure there are several secondary messengers, such as Ca, H2O2 and NO (Atkinson et al., 1990, Zhang et al., 2001, Neill et al., 2002, Garcia-Mata and Lamattina, 2009) that contribute to the stomatal closure. Passive loss of turgor pressure also results in stomatal closure.
Since stomatal closure has negative effects on CO2 uptake, photosynthesis, transpirational cooling as well as water and nutrient uptake it is important to close the stomata only when the benefit of water retention outweighs the negative effects. To be able to close the stomata during unfavourable conditions there are several mechanisms and signalling pathways leading to stomatal closure. These pathways can be divided into hydro passive and active stomatal closure.
3.1.1. Hydro passive stomatal closure
Hydro passive stomatal closure occurs when the water evaporation from the guard cells is too low to be balanced by water movement into these cells. The water content in the cells is then rapidly reduced to the extent where the osmotic pressure is reduced and the cells lose turgor pressure and shrink (Luan, 2002). When this happens the guard cells are unable to maintain the shape and the stomatal pore is covered.
Some studies have shown that passive stomatal closure is important in ferns and Lycopods, but not in Angiosperms and Gymnosperms (Franks and Farquhar, 2007, Brodribb and McAdam, 2011). This is because in Angiosperms and Gymnosperms the guard cells closely interact with their subsidiary cells. When the guard cells lose turgor pressure the subsidiary cells also lose turgor pressure and the force from the subsidiary cells pulls the guard cells apart, opening the stomata. This hydro passive opening is called the “wrong-way” response (Franks and Farquhar, 2007). In contrast the guard cells of ferns and Lycopods do not interact closely with their subsidiary cells.
The loss of turgor pressure in the subsidiary cells in these plants does therefore not result in the guard cells being pulled apart. The simultaneous loss of turgor in the guard cells will in these plants be enough to close the stomata.
Hydro passive and active stomatal closure pathways: Active stomatal closure: https://www.intechopen.com
ABA as well as elevated levels of CO2 activates signalling pathways leading to stomatal closure (Kim et al., 2010). ABA is produced in the roots and leaves during water stress and is transported to the guard cells. ABA is transported into the guard cells by ATP-binding cassette (ABC) transporters that are located in the plasma membrane (Kang et al., 2010). When the ABC transporters are knocked out the ABA uptake is lower, stomata remain more open during drought and the stress tolerance is decreased (Kang et al., 2010). The ABA signals are first recognized by several receptors. PYR/PYL/RCAR (PYRABACTIN RESISTANCE/ PYRABACTIN RESISTANCE –LIKE/REGULATORY COMPONENT OF ABA RESPONCE) proteins have been shown to function as ABA receptors (Klingler et al., 2010). Another protein GCR2 (G protein coupled receptor) has also been shown to be a ABA receptor (Liu et al., 2007).
The size of the stomatal opening is regulated by the turgor pressure and cell volume of the guard cells (Schroeder et al., 2001, Kim et al., 2010). Regulation of stomatal opening is linked to transport of ions and water through channel proteins across the plasma and vacuole membrane (Kim et al., 2010). ABA induces the production of reactive oxygen species (e.g. H2O2), which in turn acts as a trigger for NO production, inhibition of membrane proton pumps and Ca influx across both the plasma and vacuole membranes. H-ATPases that are hyperpolarizing the plasma membranes must be inhibited to induce ABA mediated stomatal closure (Merlot et al., 2007). The increased Ca levels activate slow and rapid type anion channels, generating an anion efflux from the cells. The anion efflux depolarizes the membrane, which in turn causes K efflux through K out channels across both the vacuole and the plasma membrane. Simultaneously Ca2+ also inhibits K in channels (Wasilewska et al., 2008). Malate is also converted to starch reducing the osmotic potential and turgor pressure further (Kim et al., 2010). The plasma membrane is thus depolarized, the turgor pressure and cell volume reduced and the stomata close (Kim et al., 2010).
Plant Regulators: Insensitivity is in the genes; Christopher D. Rock [Author Vitae], Ralph S. Quatrano)1
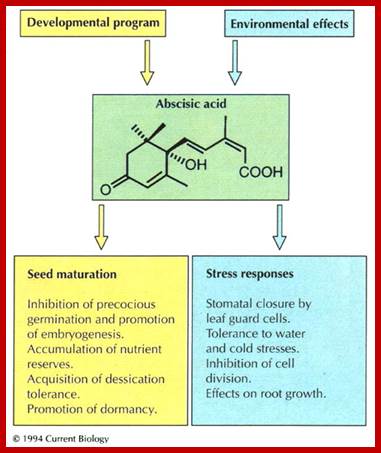
Plant Regulators: Insensitivity is in the genes
Christopher D. Rock, Ralph S. Quatrano; ;http://www.cell.com
Cell signaling in plant pattern formation
Plant cells are not able to change their position and orientation, because they are interconnected by rigid cell walls. Therefore, formation of correct tissue patterns relies on a mechanism that determine the fate of each cell depending on its position. What makes such developmental process possible?
For the cells to know their positions in organ primordia, they need to communicate each other. Cells exchange "positional cues" to decide their cell fate. The theory of position-dependent cell fate determination is the basic principle of plant development.
Cell-cell communication is usually mediated by ligand-receptor pathways, which are seen in both plants and animals. However, recent studies revealed the existence of a plant-specific cell-cell communication mechanism.
It has been known that plant cells a connected by small tunnels across the cell walls, termed the plasmodesmata (PD). Since PDs serve as a channel for plant viral spread, PDs have been studied mostly by plant pathologists. However, as PDs became recognized as essential channels to transmit developmental signals, developmental biologists are now interested in the regulation of PD functions.
http://bsw3.naist.jp